install.packages(c("devtools", "RColorBrewer"))
devtools::install_gitlab('jbferet/prosail')8 Simulating spectra
R packages
A package is a collection of previously programmed functions, often including functions for specific tasks. It is tempting to call this a library, but the R community refers to it as a package.
There are two types of packages: those that come with the base installation of R and packages that you must manually download and install using install.packages("package"). You can load a package with library(package).
To work with the tutorial, first install the following packages.
You can keep your packages up to date with with the following command:
update.packages()Important, you only need to install a package once (and never in knitr), but you need to load it every time you start a new R-Session! (RColorBrewer is for my plotting function.)
Load the prosail package for manipulating and simulating hyperspectral data (Féret and de Boissieu, 2024). The package contains the PROSPECT and PROSAIL model.
library(prospect)
library(prosail)
library(RColorBrewer) # for plotting color optionsNamespace
Sometimes functions from different packages have a common name (e.g., filter() in the stats and dplyr packages). We therefore need to tell R which function it should use by defining the namespace for the function:
stats::select()
dplyr::select()PROSPECT
PROSPECT is a widely used leaf reflectance (or transmittance) model which simulates reflectance values between 400 and 2500 nm (Féret et al., 2017; Jacquemoud et al., 2009; Jacquemoud and Baret, 1990).
PROSPECT requires a set of parameters describing structure and chemical composition of leafs. See help by typing ?PROSPECT.
There are default values for each parameter. These default values were included with the intention to provide an easy access to the model and should be used with care in any scientific approach! Calling PROSPECT() returns a dataframe.
spec <- PROSPECT(N = 1.5, CHL = 40, CAR = 8, ANT = 1.0, BROWN = 0.0,
EWT = 0.01)
head(spec) wvl Reflectance Transmittance
1 400 0.04312061 0.0003485836
2 401 0.04311894 0.0003382251
3 402 0.04311758 0.0003297392
4 403 0.04311690 0.0003254520
5 404 0.04311715 0.0003270545
6 405 0.04311859 0.0003360348Now, let’s jump into the simulation of reflectance values. Simulate two spectra with Chlorophyll content: 1) with 5 \(\mu g/cm^2\) and 2) 90 \(\mu g/cm^2\). Plot the two spectra and describe the difference.
## Simulate first spectrum with lower chlorophyll content
spec1 <- PROSPECT(N = 1.3, CHL = 5, CAR = 10, BROWN = 0, EWT = 0.01)
## Simulate second spectrum with higher chlorophyll content
spec2 <- PROSPECT(N = 1.3, CHL = 90, CAR = 10, BROWN = 0, EWT = 0.01)
# plot spectrum
plot(spec1$wvl, spec1$Reflectance, col=2, ylim = c(0,0.5), type="l",
xlab="Wavelength", ylab="Reflectance")
lines(spec2$wvl, spec2$Reflectance, col=4, new = FALSE)
legend("topright",
legend=c(expression(paste('CHL = 5 ', mu*g*cm^-2)),
expression(paste('CHL = 90 ', mu*g*cm^-2))),
lty=1, col=c(2,4), bty='n')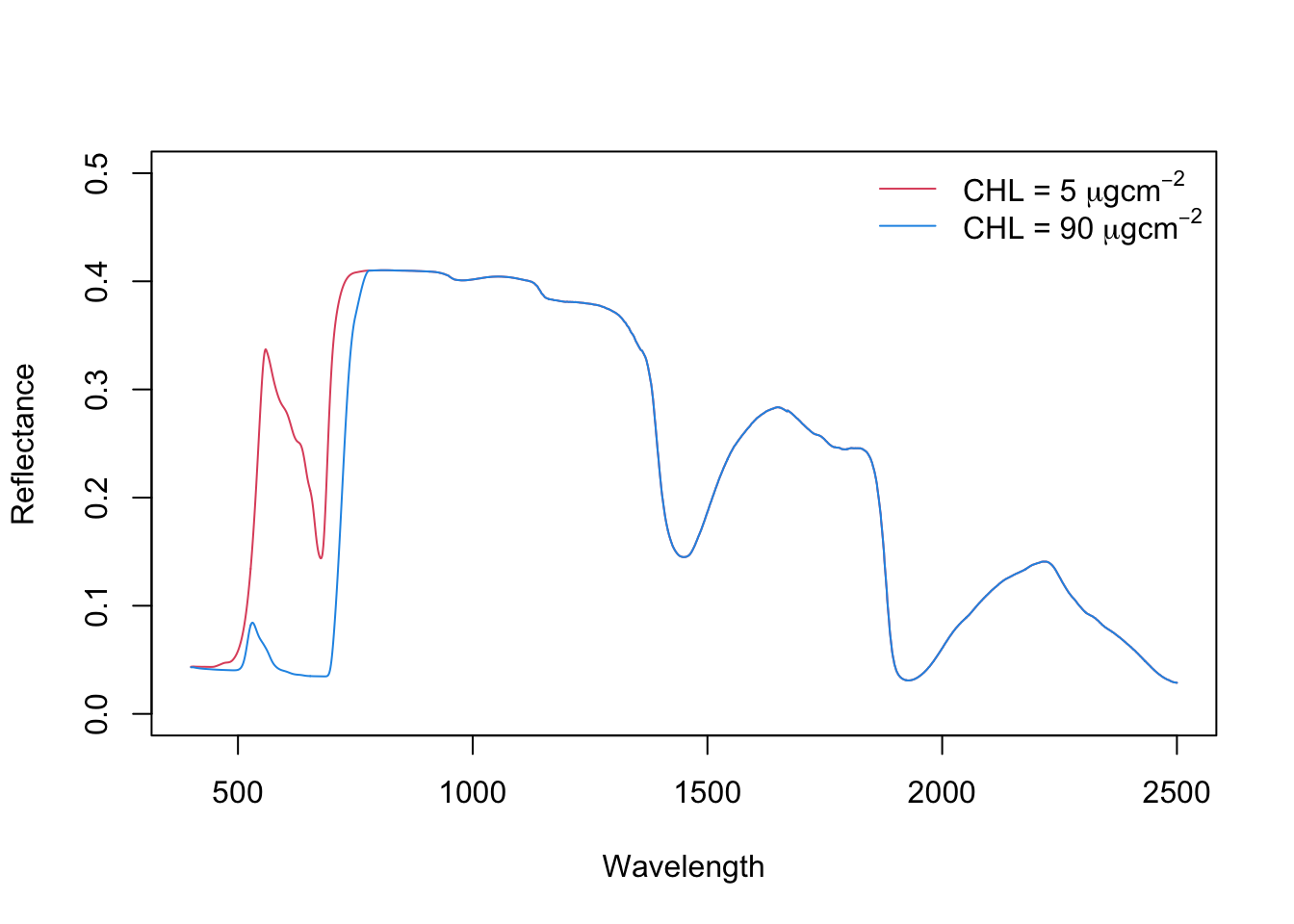
Increasing chlorophyll content causes a) absorption in blue and red spectrum and b) red-shift of red-edge (to the right).
PROSAIL
PROSAIL is a fusion of PROSPECT (leaf reflectance and transmittance) and SAIL (plant canopy reflectance). Thus, the model simulates canopy reflectance based on leaf biochemical and structural parameters as well as canopy and observation geometry. PROSAIL been used for the last thirty years to simulate the spectral and directional reflectance of plant canopies in the solar domain. It links the spectral dimension of the reflectance, which is mainly related to the biochemical content of the leaves, to the directional dimension, which is mainly related to the architecture of the canopy (Jacquemoud et al., 2009).
?PRO4SAILThe PRO4SAIL function of the prosail package runs the PROSAIL model. Check out the help pages for a description of the function arguments and model parameters. Here is an example of selected parameters:
| Model | Parameter | Description |
|---|---|---|
| PROSPECT | N | Leaf structure parameter |
| CHL | Cholorophyll a + b content | |
| EWT | Equivalent water thickness | |
| ANT | Anthocyanin content | |
| BROWN | Brown pigments content | |
| LMA | Leaf Mass per Area | |
| SAIL | lai | Leaf area index |
| TypeLidf | Leaf inclination distribution function | |
| LIDFa | if TypeLidf = 1, controls the average leaf slope if TypeLidf = 2, controls the average leaf angle | |
| LIDFb | if TypeLidf = 1, controls the distribution’s bimodality if TypeLidf = 2, unused | |
| q | Hot spot parameter | |
| tts | Solar zenith angle | |
| tto | Viewing zenith angle | |
| psi | Azimuth Sun / Observer | |
| rsoil | Soil reflectance |
The PRO4SAIL function requires you to define the leaf properties (Input_PROSPECT), soil reflectance (predefined for wet and dry soil), and canopy parameters. The example below, defines the leaf parameters and uses a reflectance profile from a wet soil.
# define input variables for PROSPECT.
Input_PROSPECT = data.frame('CHL' = 40, 'CAR' = 8, 'ANT' = 0.0,
'EWT' = 0.01, 'N' = 1.5)
# wavelength
wvl <- SpecPROSPECT_FullRange$lambda
# soil reflectance
rsoil <- SpecSOIL$Wet_Soil
# run PROSAIL with 4SAIL
Ref_4SAIL <- prosail::PRO4SAIL(Input_PROSPECT = Input_PROSPECT,
TypeLidf = 1, LIDFa = -0.35, LIDFb = -0.15,
lai = 5, q = 0.01, tts = 30, tto = 10, psi = 0,
rsoil = rsoil)PRO4SAIL returns a list containing different versions of reflectance factors. In the context of satellite measurements, we are interested in the bi-directional reflectance factor. So, we extract rsot and pair it with wavelength in a new data frame (here called bf).
rddt: bi-hemispherical reflectance factorrsdt: directional-hemispherical reflectance factor for solar incident fluxrdot: hemispherical-directional reflectance factor in viewing directionrsot: bi-directional reflectance factorfcover: Fraction of green Vegetation Cover (= 1 - beam transmittance in the target-view path)abs_dir: canopy absorptance for direct solar incident fluxabs_hem: canopy absorptance for hemispherical diffuse incident fluxrsdstar: contribution of direct solar incident flux to albedorddstar: contribution of hemispherical diffuse incident flux to albedo
bf <- data.frame(wvl = wvl, refl = Ref_4SAIL$rsot)
plot(refl ~ wvl, data=bf, type='l')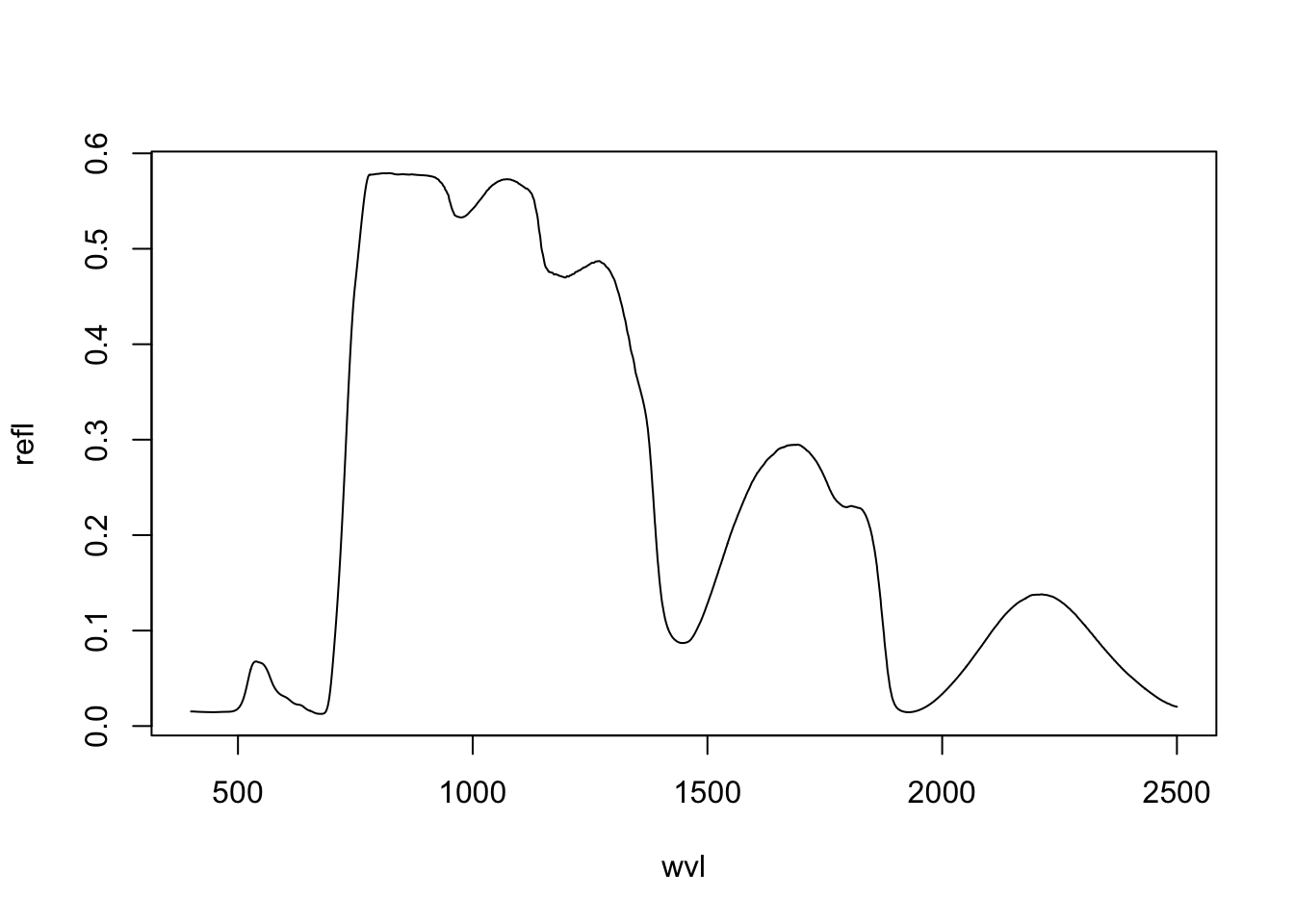
To test a range of different parameters, you need to run the PRO4SAIL function multiple times. For example, if you we want to test the effect of LAI on reflectance values, you can simulate the associated spectra by repeatedly calling PRO4SAIL and changing the input argument each time. This is quite cumbersome. So, for the purpose of this exercise, I created two convenience functions: 1) PRO4SAIL_batch and 2) PRO4SAIL_plot. To use the two functions in your assignment, you need to include (copy/paste) them into your markdown documents.
Code
PRO4SAIL_batch <- function(...) {
arg <- list(...)
Input_PROSPECT <- data.frame('CHL' = 40, 'CAR' = 8, 'ANT' = 0.0,
'EWT' = 0.01, 'LMA' = 0.009, 'N' = 1.5)
varname <- names(arg)[1]
values <- arg[[1]]
wvl <- SpecPROSPECT_FullRange$lambda # wavelength
result <- data.frame()
for (val in values) {
l <- list(Input_PROSPECT=Input_PROSPECT, var=val, TypeLidf=1,
LIDFa=-0.35, LIDFb=-0.15, rsoil = SpecSOIL$Wet_Soil)
names(l)[2] <- varname
tmp <- data.frame(wvl = wvl, refl = do.call(prosail::PRO4SAIL, l)$rsot)
if (!all(is.na(tmp$refl))) {
tmp[, varname] <- val
result <- rbind(result, tmp)
}
}
return(result)
}
PRO4SAIL_plot <- function(spectra, ylim=c(0, 0.5)) {
require(RColorBrewer)
varname <- names(spectra)[3]
values <- sort(unique(spectra[, 3]))
colours <- brewer.pal(length(values), 'Spectral')
for (i in seq_along(values)) {
if (i == 1) {
plot(spectra$wvl[spectra[, varname] == values[i]], spectra$refl[spectra[, varname] == values[i]],
ylim = ylim, col = colours[i], type='l', xlab="Wavelength [nm]",
ylab = "Reflectance")
} else {
lines(spectra$wvl[spectra[, varname] == values[i]],
spectra$refl[spectra[, varname] == values[i]], col = colours[i])
}
}
labels <- paste(varname, "=", values)
legend('topright', legend=labels, col=colours, lty=1, bty='n')
}LAI
With PRO4SAIL_batch you can simulate bi-directional reflectance by changing a single SAIL parameter. For example, below we simulate reflectance LAI values 1 to 6. If you want to simulate the reflectance for a different parameter, just change the argument name and values in the function call.
spectra.lai <- PRO4SAIL_batch(lai=1:6)With PRO4SAIL_plot you can visualize the output.
PRO4SAIL_plot(spectra.lai, ylim=c(0, 0.5))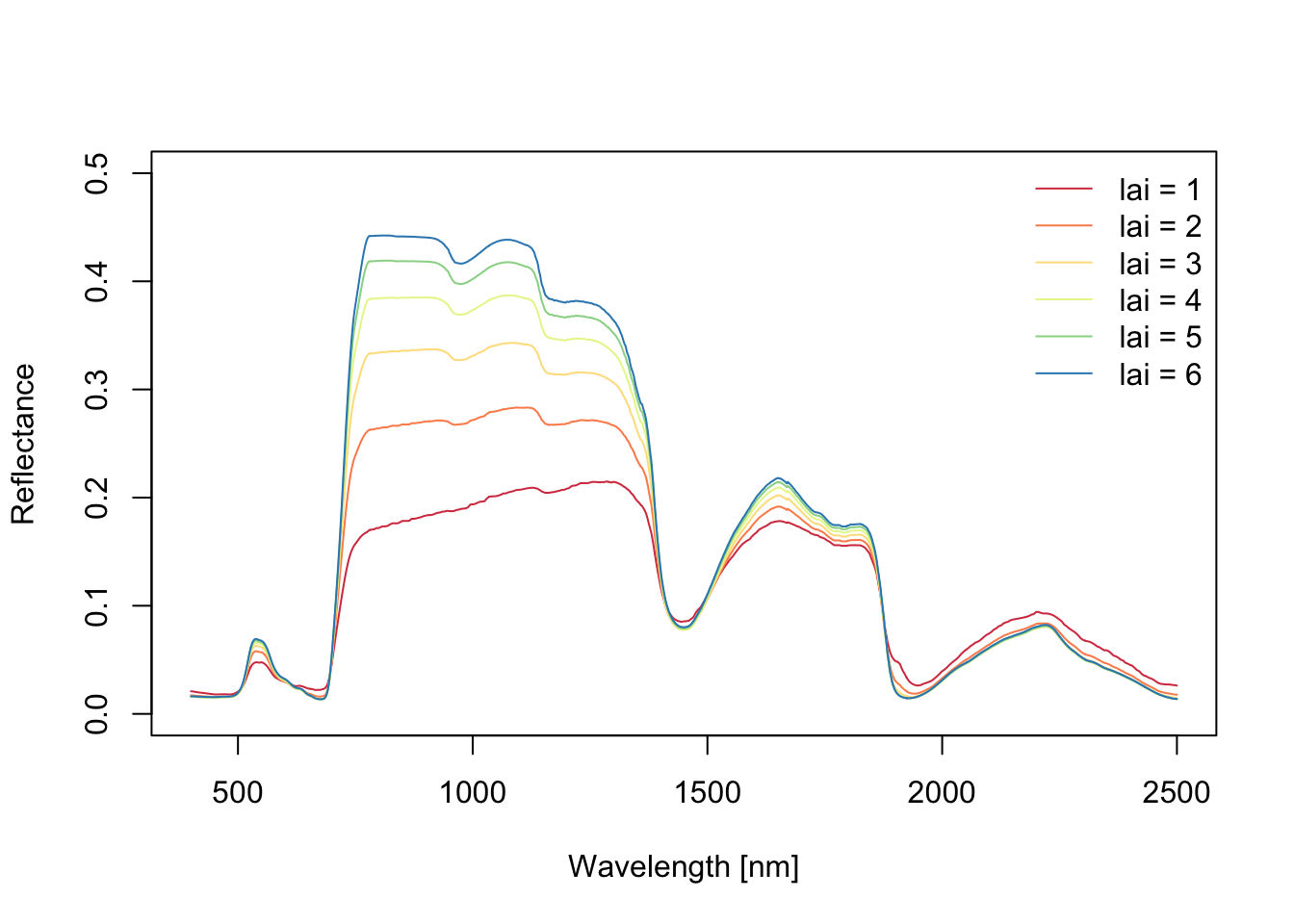
Increasing LAI causes: a) increasing NIR, b) decreasing reflectance in red, and c) comparably small impact on shortwave-infrared.